Main content start
The future of proteins
An expert in protein structure and function explains how rapidly evolving study of these complex molecules is revealing ever-greater understanding of human life.
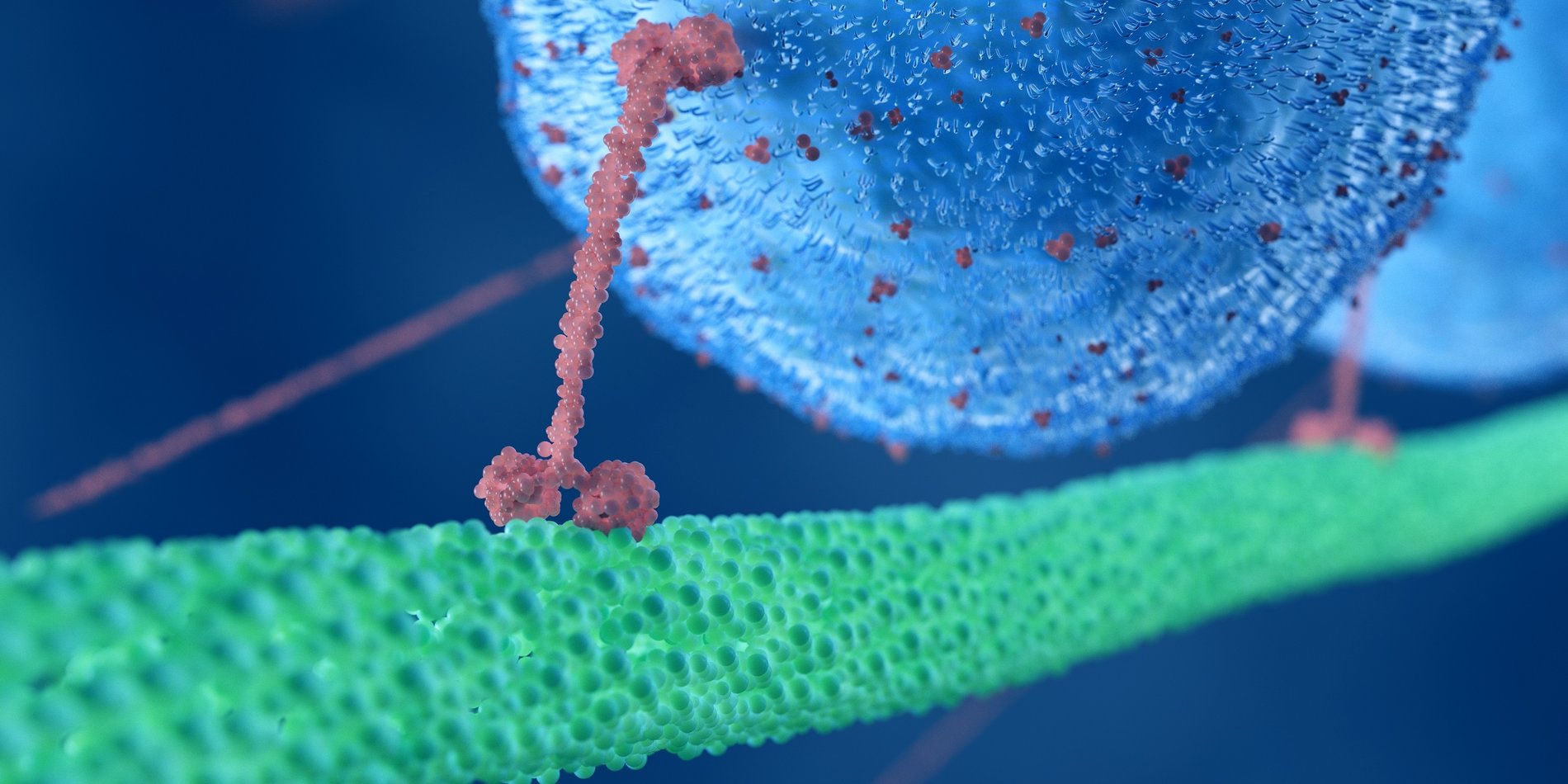
While DNA may be the blueprint of life, proteins are the workhorses, says Polly Fordyce, a bioengineer, explaining how one of her favorites, kinesin, “walks” in 8-nanometer steps transporting chemical cargo through the body.
More remarkable still, Fordyce says, kinesin is just one among thousands of “incredible” proteins that make life happen, as she tells host Russ Altman on this episode of Stanford Engineering’s The Future of Everything podcast.